ISOLATION, CHARACTERIZATION AND PATHOGENIC POTENTIAL OF FUNGI FROM WASHING MACHINES
DOI:
https://doi.org/10.33003/fjs-2026-1003-4329Keywords:
Washing machine, antifungal, Resistance, Biofilm, EnzymesAbstract
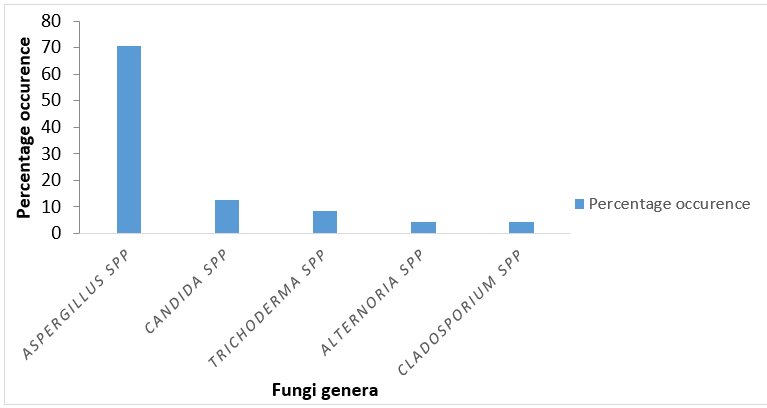
This study aimed to profile the antifungal resistance pattern of fungi isolated from washing machines sampled from hospitals, hotels, student hostels, homes and laundry services shops in Ogbomoso. Isolation procedures for collected samples were carried out using standard microbiological techniques. Fungal isolates were subjected to antifungal drugs to determine their susceptibility pattern. Isolates with multiple resistance ≥ 60 % were selected for molecular characterization. The ability of the selected isolates to produce biofilm was assessed and quantified at 492nm and toxin production was also determined. The potential of the fungi to produce enzymes such as protease, amylase, cellulase, pectinase and collagenase were screened. Questionnaires were given to the staff of hotels and laundry service shops on the treatment of washing machines. A total of 24 fungal isolates belonging to 5 genera of Aspergillus, Trichoderma, Candida, Cladosporium and Alternaria. Aspergillus species were found to be the predominant (70.8%). The highest resistance to anti-fungal drugs employed was from Aspergillus niger with 80% resistant. Aspergillus niger was the only biofilm producer, belonging to the weak biofilm former. Two isolates out of the selected four fungi were able to produce aflatoxins while they all have the potential to produce more than five enzymes. Both the hotels and laundry services are frequently (82%) treated and their source of water is treated once in a month as reported by 63% of the respondents. Washing machines sampled in this study were reservoirs for multi-anti-fugal drug- resistant isolates, and this could pose serious threat to public health.
References
Adeoyo, O. R. and Omolola, O. G. (2022). Evaluation of indoor and outdoor fungal flora of two poultry farms in Akungba-Akoko andAyegunle-Akoko. Saudi J. Biomed. Res. 7(3), 114–118 (2022).
Agboola, J.O., Ayandele, A.A., Amao, J.A. (2021) Isolation and Characterization of Bacteria with Multiple Drug Resistance from Pig Dung.Trop J Nat Prod Res. 5(8):1506-1514.
Ali, A., Zahra, A., Kamthan, M., Husain, F. M., Albalawi, T., Zubair, M., Alatawy, R., Abid, M. and Noorani, M. S. (2023). Microbial Biofilms: Applications, Clinical Consequences, and Alternative Therapies. Microorganisms, 11(8), 1934
Ali, J., Sohail, A., Wang, L., Rizwan Haider, M., Mulk, S., and Pan, G. (2018). Electro-microbiology as a promising approach towards renewable energy and environmental sustainability. Energies 11:1822
Amao, J.A., Omojasola, P.F., Madhumita, B. (2019). Isolation and Characterization of some Exopolysaccharide producing Bacteria from Cassava Peel Heaps. Sci Afr.; 4(e00093):1-11
Ammann, H. M. (2003). Is indoor mold contamination a threat to health? J Environ Health.66(2)47–49.
Babicˇ, M. N., Gostincˇar, C. and Gunde-Cimerman, N. (2020). Microorganisms populating the water-related indoor biome. Applied Microbiologyand Biotechnology. 104, 6443–6462.
Clinical and Laboratory Standards Institute (2010). Method for Antifungal Disk Difusion Susceptibility Testing of Nondermatophyte Filamentous Fungi; Approved Guideline. CLSI document M51-A. Wayne: Clinical and Laboratory Standards Institute.
Copeland, K. and Purvis, A.R. (2017). A Retrospective Chart Review of Chronic Wound Patients Treated with Topical Oxygen Therapy. Adv. Wound Care6, 143–152.
de Souza, M., de Oliveira, C., Annes, I., Castro S., Rocha, B. and Silva C. (2012). Antibiotic Resistance in staphylococcus species of animal origin. In Antibiotic Resistant Bacteria - A Continuous Challenge in the New Millennium, pp. 273–303.
Dirk, P., Bockmühl, J. S. and Laura, R. (2019). Laundry and textile hygiene in healthcare and beyond.Microbial Cell6(7): 299-306.
Elsheikh, E. G., Abuzied, M, A., Moalim, N.B. and Ali, A.H. (2023). Detection of fungal contamination of ward furnishing and medical equipment used in intensive care unit andneonatal intensive care unit. J. Drug Deliv. Ther. 13(3), 47–50.
Irankhah, S., Abdi Ali, A., Mallavarapu, M., Soudi, M. R., Subashchandrabose, S. and Gharavi, S. (2019). Ecological role of Acinetobacter calcoaceticus GSN3 in natural biofilm formation and its advantages in bioremediation. Biofouling 35, 377–391
Jacksch, S., Zohra, H., Weide, M., Schnell, S. and Egert, M. (2021). Cultivation-Based Quantification and Identification of Bacteria at Two Hygienic Key Sides of Domestic Washing Machines. Microorganisms 9, 905
Johani, K., Malone, M., Jensen, S., Gosbell, I., Dickson, H., Hu, H. and Vickery, K. (2017). Microscopy Visualisation Confirms Multi-Species Biofilms Are Ubiquitous in Diabetic Foot Ulcers. Int. Wound J. 14, 1160–1169.
Lebeaux, D., Ghigo, J.-M. and Beloin, C. (2014). Biofilm-Related Infections: Bridging the Gap between Clinical Management and Fundamental Aspects of Recalcitrance toward Antibiotics. Microbiol. Mol. Biol. Rev. 78, 510–543
Makovcova, J., Babak, V., Kulich, P., Masek, J., Slany, M., Cincarova, L. (2017). Dynamics of mono- and dual-species biofilm formation and interactions between Staphylococcus aureus and gramnegative bacteria. Microbial Biotechnology. 10, 819–832.
Marchese, S., Polo, A., Ariano, A., et al. (2018). Aflatoxin B1 and M1: biological properties and their involvement in cancer development. Toxins (Basel).10(6):214.
Muhammad, M. H., Idris, A. L., Fan, X., Guo, Y., Yu, Y., Jin, X., Qiu, J., Guan, X. and Huang, T. (2020). Beyond Risk: Bacterial Biofilms and Their Regulating Approaches. Front. Microbiol. 11:928.
Nix, I. D., Frontzek, A. and Bockmühl, D. P. (2015). Characterization of microbial communities in household washing machines. Ten side Surfactants Deterg. 52(6), 432–440.
Parcell, B. J., Oravcova, K., Pinheiro, M. et al. (2018). Pseudomonas aeruginosa intensive care unit outbreak: winnowing of transmissions with molecular and genomic typing. Journal of Hospital Infections. 98(3):282–288.
Raed, N. A., Mohammed, H. K. and Basil, A. (2016).Abbas Microbiology Department, College of Veterinary Medicine, Uni.Of Basrah, Iraq. January 2016; Accepted 9 March 2016
Rainer, S., Alexander, B., Felice, A. and Mauro, C. (2020). Durability of washing machines under real life conditions: Definition and application of a testing procedure. J Clean Prod. 10; 261: 121222.
Rawson, T. M., Fatania, N. and Abdolrasouli, A. (2022). UK standards for microbiology investigations of ear infection (SMI B1) are inadequate for the recovery of fungal pathogens and laboratory diagnosis of otomycosis: A real-life prospective evaluation. Mycoses 65(4), 490–495.
Shemse, S., Woldaregay, E. A., Aminu, S., Tewachew, A., Zelalem, D., Wude, M., Adane, M. and Tamrat, A. (2020). Bacterial Profiles and Antimicrobial Susceptibility Pattern of Isolates from Inanimate Hospital Environments at Tikur Anbessa Specialized Teaching Hospital, Addis Ababa, Ethiopia. Infection and Drug Resistance13 4439–4448
Stapleton, K., Hill, K., Day, K., Perry, J. D. and Dean, J. R. (2013). The potential impact of washing machines on laundry malodor generation. Lett Appl Microbiol 56:299–306
Teufel, L., Pipal, A., Schuster, K. C., Staudinger, T. and Redl, B. (2010). Material dependent growth of human skin bacteria on textiles investigated using challenge tests and DNA genotyping. Journal of Applied Microbiology. 108, 450–461.
Tischner, Z., Kredics , L., Marik , T. Vörös, K., Kriszt, B., Péter, B. and Magyar, D. (2019). Environmental characteristics and taxonomy of microscopic fungi isolated from washing machines. Fungal Biol.123(9), 650–659.
Vasudevan, R. (2014). Biofilms: microbial cities of scientific significance. Journal of Microbiology. 1 (3),00014.
Whitehead, K., Eppinger, J., Srinivasan, V., Khalid Ijaz, M., Nims, R.W. and McKinney, J. (2022). Potential for microbial cross contamination of laundry from public washing machines. Microbiol. Res. 13(4),995–1006.
WHO. (2014). Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva, Switzerland.
Zhang, M., Han, W., Gu, J., Qiu, C., Jiang, Q., Dong, J., Lei, L. and Li, F. (2022). Recent advances on the regulation of bacterial biofilm formation by herbal medicines. Front. Microbiol. 13:1039297.
Zupancicˇ, J., Raghupathi, P. K., Houf, K., Burmølle, M., Sørensen, S.J. and Gunde-Cimerman, N. (2018). Synergistic Interactions in Microbial Biofilms Facilitate the Establishment of Opportunistic Pathogenic Fungi in Household Dishwashers. Frontier of Microbiology. 9, 21.
Fox, E.P., Biu, C.K., Nett, J.E., Hartzen, A.R., Andes, D.R. and Hogan, D.A. (2014). The Candida albicans pescadillo homolog is required for normal hypha-to-yeast morphogensesis and tolerance to antifungal drugs. PLOS pathogens, 10 (10), e1004450
Schaller, M., Borelli, C., Korting, H.C. and Hube, B. (2018). Hydrolytic enzymes as virulence factors of Candida albicans. Mycoses, 61(5), 281--291
Marin, S., Ramos, A.J., Cano-Sancho, G., Sanchis, V. (2013). Mycotoxins: Occurrence, Toxicology and exposure assessment. Food chemistry and toxicology 60: 218-237.
Downloads
Published
Issue
Section
Categories
License
Copyright (c) 2026 IFEOLUWA OMOTOLA OMOTOSO, OPEYEMI GLORY TIAMIYU, ABIODUN A. AYANDELE, JOHN AYO AMAO

This work is licensed under a Creative Commons Attribution 4.0 International License.




